Bluesky
Recent posts from @bioinfo.se,arneelof.
- No cached Bluesky posts yet. Run
make feedsand rebuild.
The Elofsson Lab develops computational methods to understand how proteins fold, interact, evolve, and function, combining machine learning, structural bioinformatics, and large-scale biological data.
Major support from KAW, DDLS, EU and Vetenskapsradet (VR).
Latest social updates from Bluesky.
The group combines structural bioinformatics, machine learning, and evolutionary analysis to understand proteins, complexes, membranes, and molecular systems.
From consensus and quality assessment methods to AlphaFold-era modelling of protein interactions and higher-order assemblies.
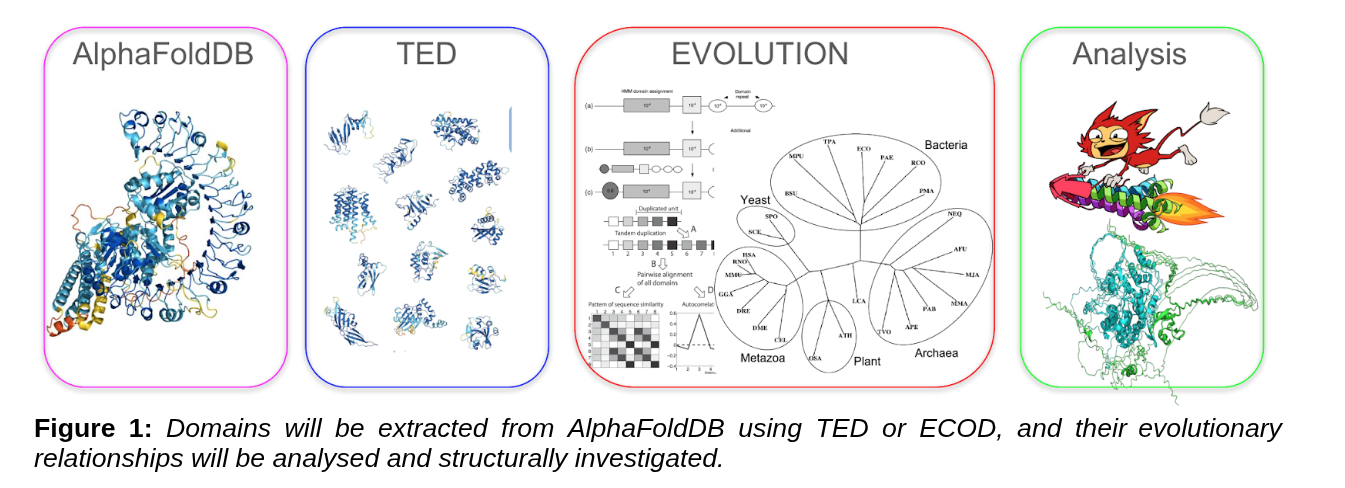
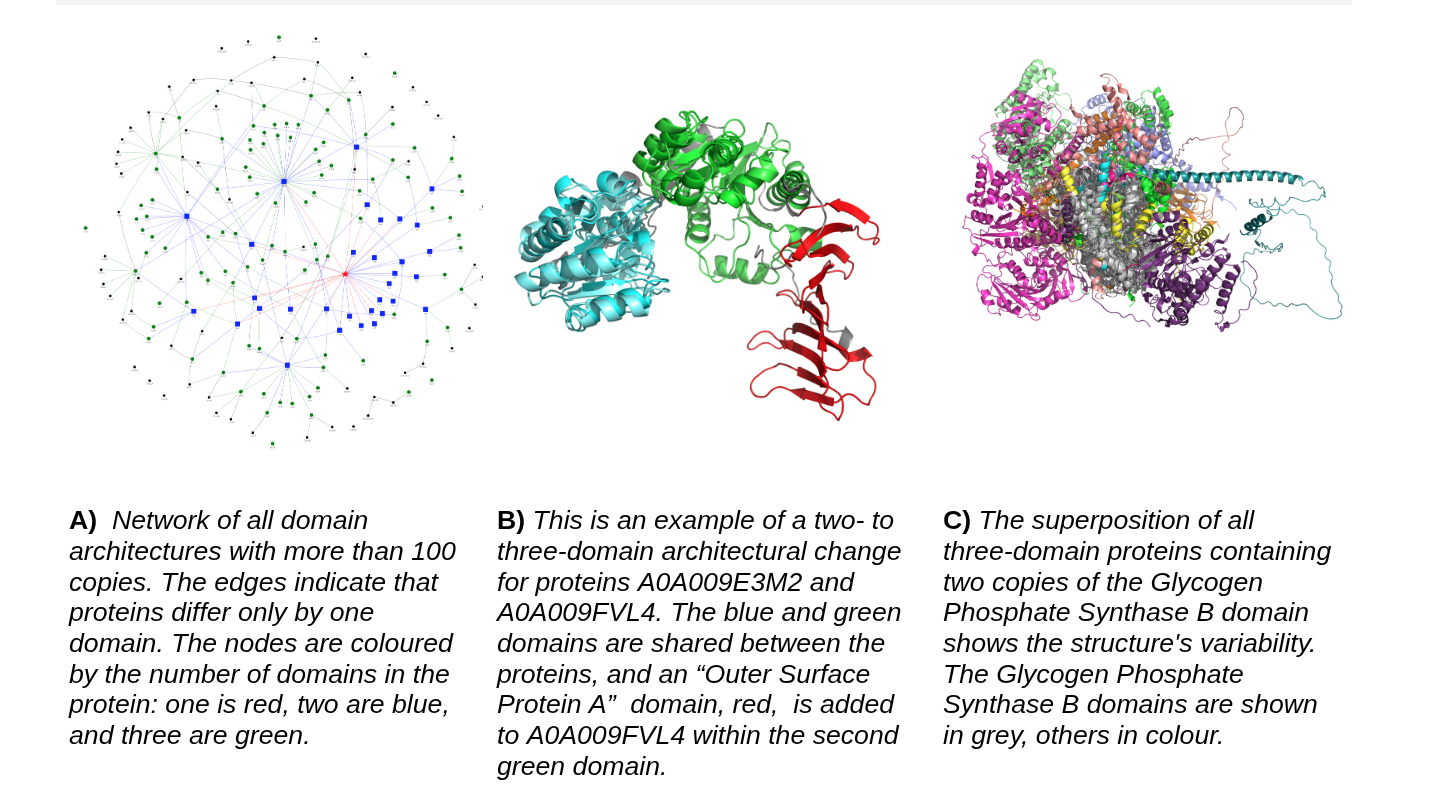
How domains change, duplicate, and rearrange across the tree of life, now with structure-aware analysis using AlphaFoldDB, TED, and ECOD.
Methods and mechanistic insight for membrane protein topology, transporters, and membrane-protein structure.
Representation learning, graph methods, and protein language models for proteoforms, interactions, and design.
Recent projects combine large-scale structural data, evolutionary analysis, and deep learning to address questions that range from molecular systems in the cell to the emergence of protein architectures across the tree of life.

The lab led by Arne Elofsson is known for longstanding contributions in protein structure prediction, membrane protein topology, protein model quality assessment, and protein evolution. Today, this foundation is being extended into AlphaFold-era modelling, proteome-scale interaction and complex prediction, and AI-driven analysis of proteoforms and molecular systems.
We offer projects at the interface of AI, protein science, and structural biology, with strong links to Stockholm University, SciLifeLab, and national computational infrastructure.