Evolution of protein domain architectures in the AI era
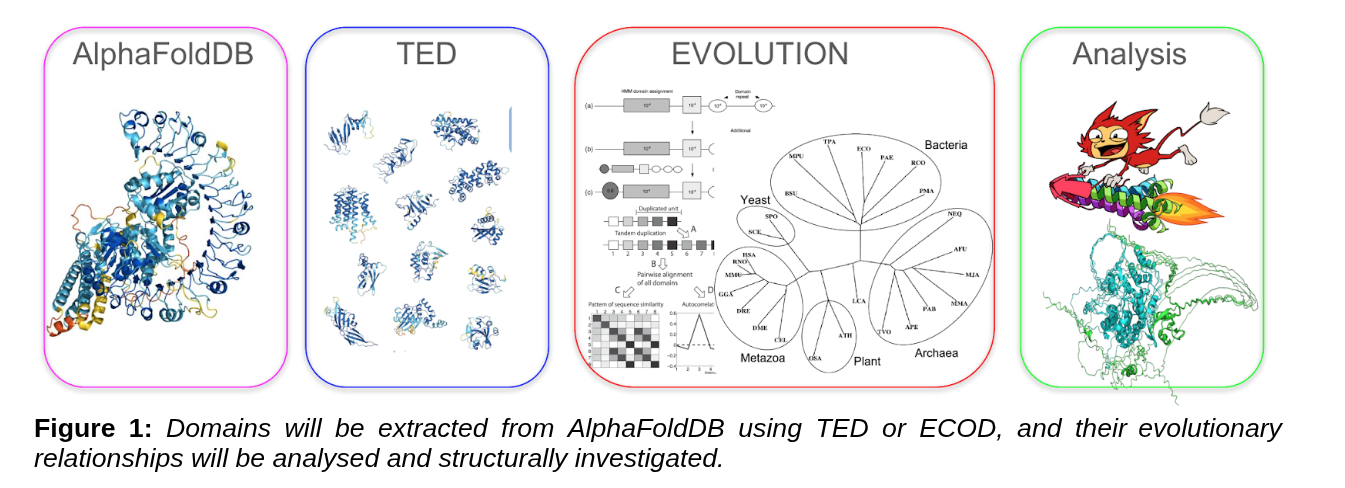
Structure-aware analysis of domain rearrangements and structural change using AlphaFoldDB, TED, and ECOD.
Amount: 4.2 MSEK
Role: Principal Investigator
Current funded projects and strategic initiatives shaping the lab's research agenda.
Structure-aware analysis of domain rearrangements and structural change using AlphaFoldDB, TED, and ECOD.
Amount: 4.2 MSEK
Role: Principal Investigator
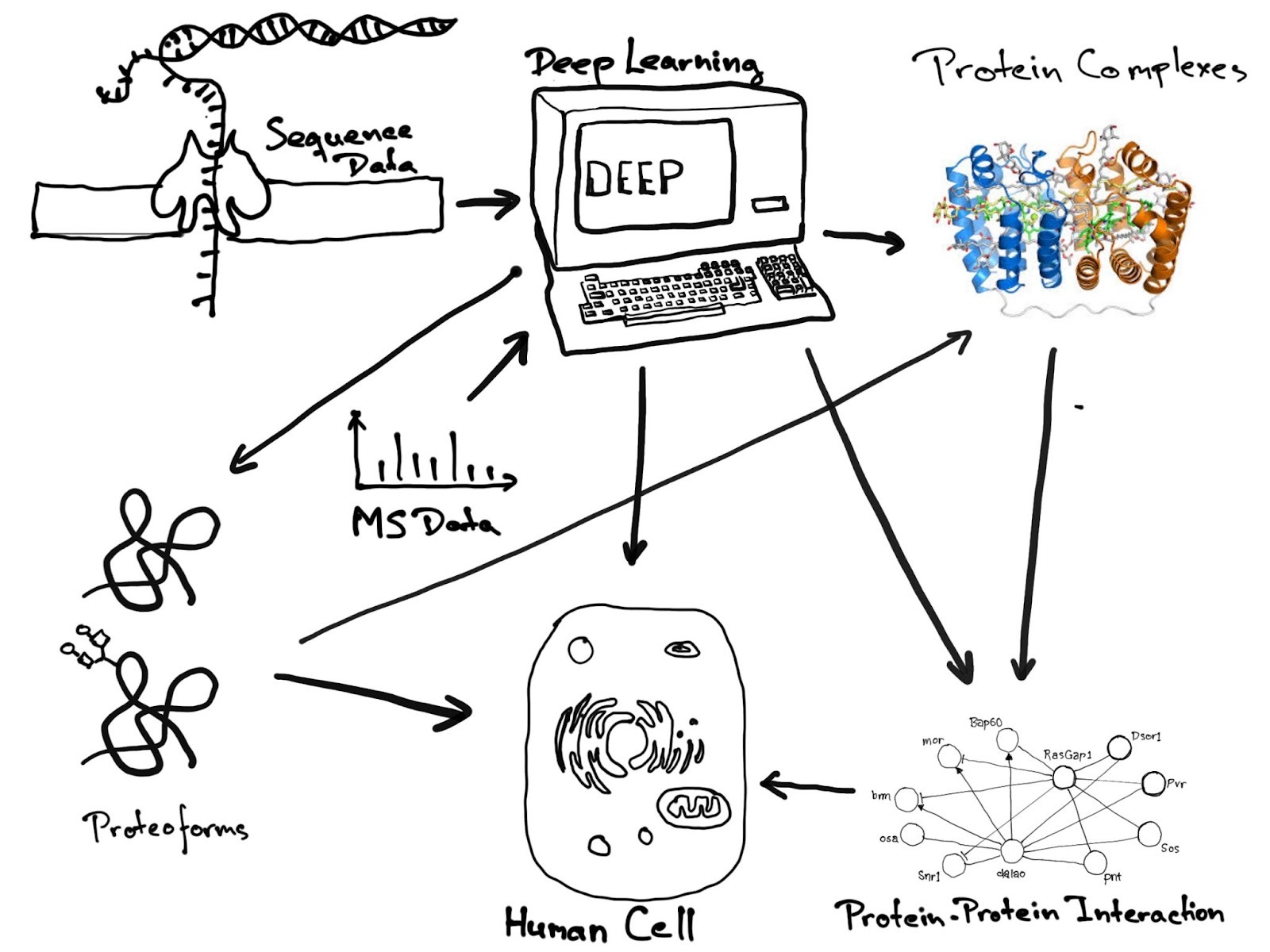
Deep-learning methods for proteoforms, protein-protein interactions, and large protein complexes.
Amount: 30 MSEK
Role: Principal Investigator
Method and infrastructure development for computational protein science.
Amount: 2.5 MSEK
Role: Principal Investigator
Center for Modelling Life at the Membrane with AI, where Arne Elofsson is co-director and co-leads the theme on comparative and predictive modelling of diverse cell membranes.
Amount: 28 MSEK
Role: Co-Principal Investigator
Collaboration with Cytiva on AI-based development of protein binders.
Amount: 2.3 MSEK
Role: Principal Investigator
Evolution of protein domain architectures in the AI-era.
Amount: 2.5 MSEK
Role: Principal Investigator
Research school for PhD students in medical bioinformatics
Amount: 30 MSEK
Role: Principal Investigator